January 8, 2026 – by Santina Russo
Still today, the impact of malaria is staggering. According to the World Health Organization’s (WHO) latest World Malaria Report, an estimated 263 million people were infected and almost 600,000 died from the disease in 2023. The burden is heaviest in African countries, which account for 94 percent of all malaria cases worldwide. But the eastern mediterranean region is also affected and has suffered 57-percent increase in incidence since 2021 with 20 in 1000 people getting the disease. Young children are especially vulnerable, as they haven’t yet developed partial immunity, and pregnant women are at risk because the infection can cause maternal anaemia, low birth weight, and infant mortality.
In a recent study published in Nature, researchers including Monica Fernández-Quintero, now in Andrew Ward’s group at the Scripps Research Institute in La Jolla, uncovered a class of antibodies capable of blocking central virulence mechanisms of Plasmodium falciparum, the parasite responsible for the most severe form of malaria. Using the CSCS supercomputer "Piz Daint", Fernández-Quintero contributed to this study with an in-depth characterization of the antibody binding interface.
A disease with profound effects
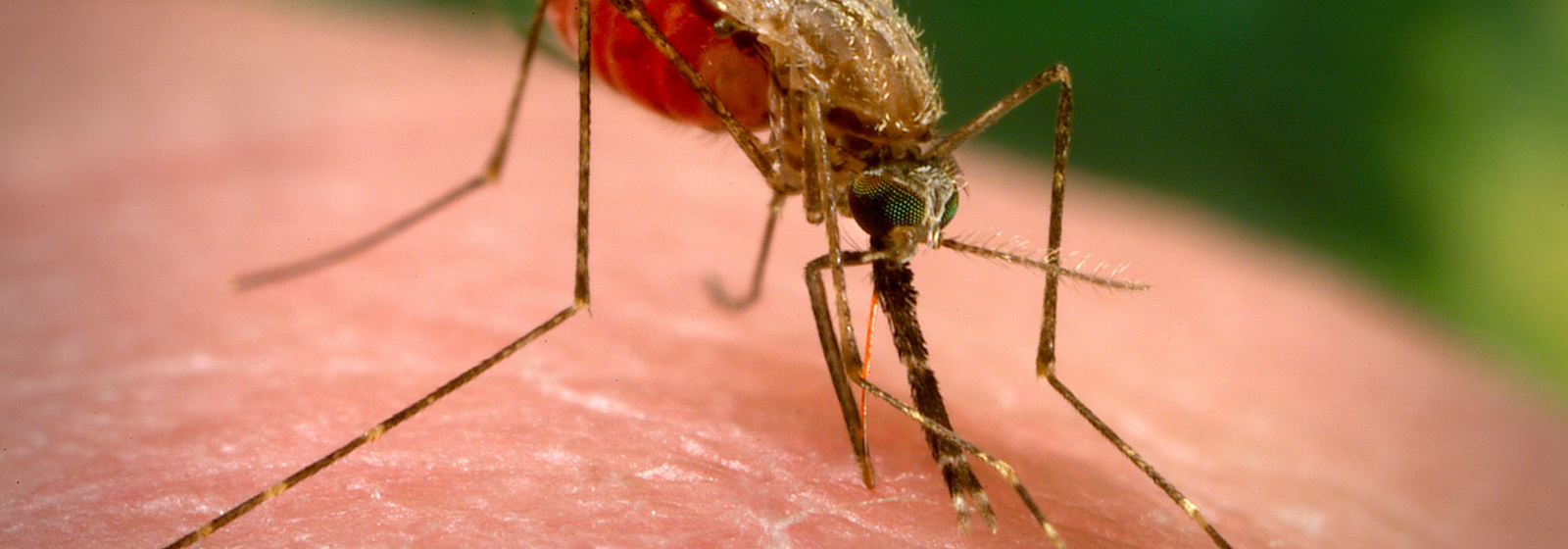
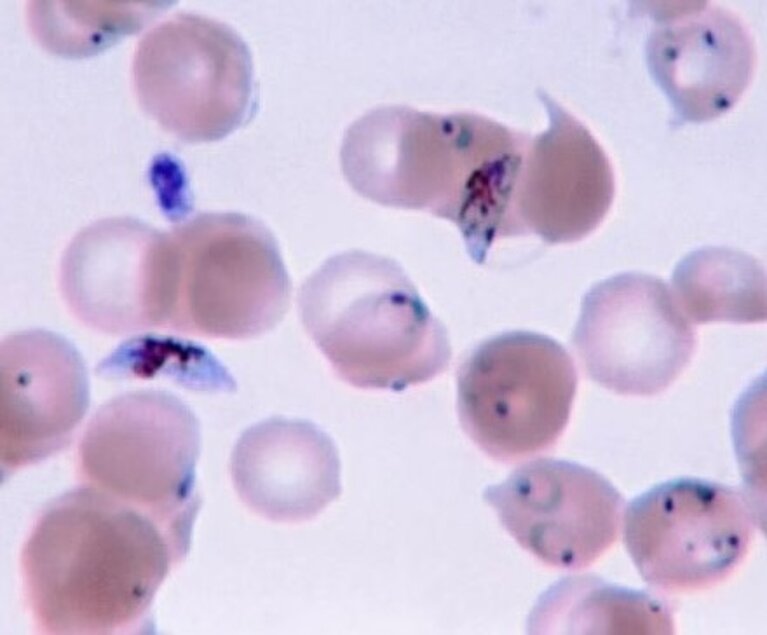
The Plasmodium falciparum parasite is transmitted by Anopheles mosquitos. When an infected mosquito bites a person, it injects the parasite’s developmental stage into the bloodstream. Once matured in the liver, the adult parasites enter the blood stream and invade red blood cells, where they multiply until they cause the cells to burst. This leads to the disease’s characteristic cyclical fevers, chills, and flu-like symptoms.
But malaria can also cause failure of vital organs, and long-term neurological effects—from cognitive deficits to motor skills impairments to epilepsy. “This underscores the importance of preventing infection, especially in young, developing brains”, said Fernández-Quintero. And the effects go beyond physical symptoms. Its high prevalence in some African countries places an immense strain on healthcare systems and economies, hindering development.
From initial to potent antibodies
Much of the disease’s damage occurs when infected red blood cells adhere to the endothelial walls of blood vessels. As the infected cells accumulate, the smallest vessels become blocked, reducing the supply of oxygen and nutrients and triggering inflammation. This clogging is mediated by the polymorphic erythrocyte membrane protein 1 (PfEMP1) of the Plasmodium parasite. In the process, it binds to receptors at the surface of endothelial cells, which form the inner lining of blood vessels.
Through extensive lab experiments and molecular dynamics simulations, Fernández-Quintero and her co-authors now identified two antibodies—isolated from two different patients—that consistently and potently inhibit the endothelial binding of five out of six PfEMP1 subclasses, thereby providing protection from the disease.
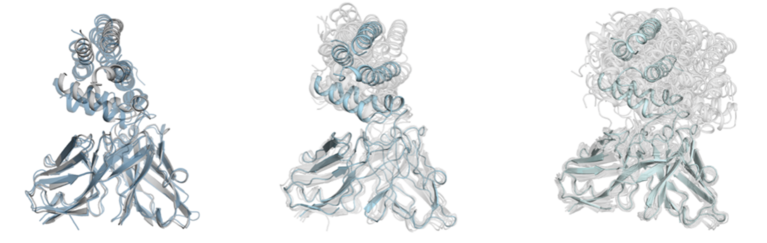
For their simulations at CSCS, the team used experimentally obtained structures of the two antibodies as the basis for modelling the antibody-antigen complexes. This term refers to antibodies bound to their target protein—in this case, a specific subdomain of the PfEMP1 protein known as CIDRα1. “This is the domain responsible for the endothelial binding that mediates the disease,” explained Fernández-Quintero. The scientists focused on the antibodies’ maturation: from the state before the infection, meaning before the immune system first encountered the virulent CIDRα1 domain, through their evolution into potent antibody against it.
A challenge: the antibodies’ diversity
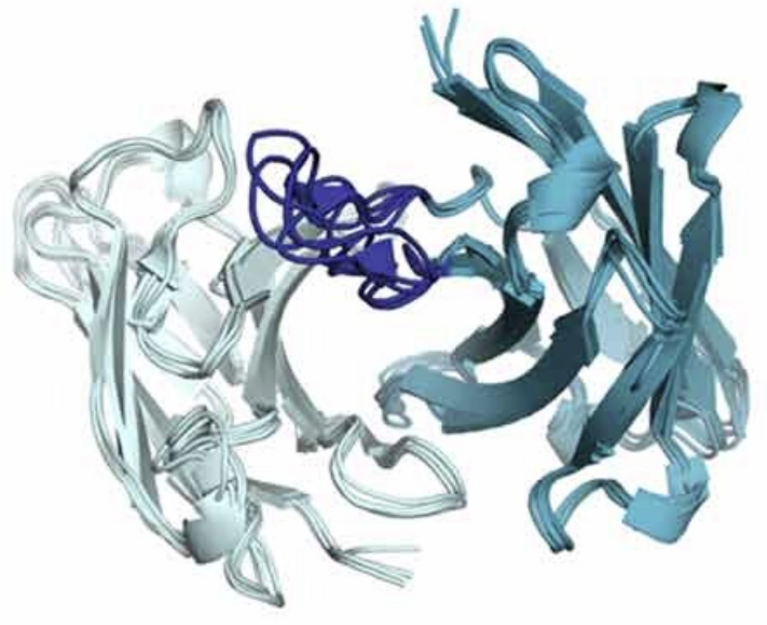
Through their simulations, Fernández-Quintero and her colleagues revealed the molecular structure of the antibody-antigen binding interface, and they analysed and quantified the structures’ flexibilities and the contacts between binding partners. The results showed that, during maturation, a rigidification of the binding interface leads to more stable interactions. This makes the mature antibodies efficient inhibitors. “With our results as a basis, these antibodies can now be further investigated as a promising option for treatment or a vaccine against malaria,” stated Fernández-Quintero.
In a separate project, the structural biologist also tested AI approaches to model antibody-antigen binding. “When it comes to predicting protein structures, antibodies pose a particular challenge,” said Fernández-Quintero. Because they evolve to respond to virulence proteins from a wide range of invading pathogens—including ones the immune system has never seen before—they are highly diverse. While a large part of their structure is almost identical across all antibodies, the region responsible for antigen binding shows a high degree of conformational diversity and is particularly hard to predict.
How to double-check AI methods
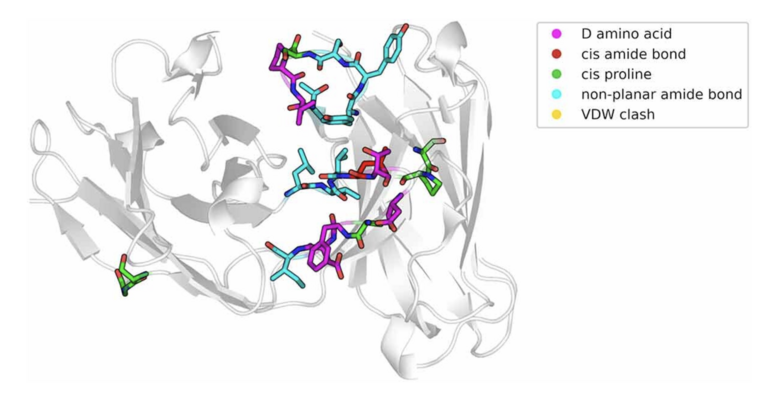
When the team compared structures resulting from different AI-enhanced antibody structure prediction tools, they discovered that many of them contained inaccuracies and errors. They found clashes between atoms, or cis-amide bonds and incorrect stereochemistry—features that are theoretically possible but never found in proteins in nature. Overall, the team examined 137 antibody models which collectively contained more than 500 physical errors.
“This can be problematic, because as a researcher using these tools you assume they work reliably and produce correct structures,” said Fernández-Quintero. Her take home message is to keep using AI methods but to check the results rigorously. “It’s essential to know how reliable current machine learning predictions really are, and what needs to be checked in the results they produce.”
To address this issue, she and her co-workers developed a tool named “TopModel” that can validate antibody structures. “After all, antibodies are among the most important classes of therapeutic proteins, and their antibody-antigen interactions are central to many of medical conditions, from malaria and other infections to different allergies and cancer,” the scientist noted. Insights into how antibodies recognize virulent proteins may therefore prove crucial for more effective treatments for a variety of diseases.
Cover Image: The Anopheles arabiensis mosquito is a known carrier of the pathogen causing malaria, the parasite Plasmodium falciparum.
(Image credit: Wikimedia Commons, CDC/James Gathany)
References:
R.A Reyes, S.S.R. Raghavan, N.K. Hurlburt et al. Broadly inhibitory antibodies to severe malaria virulence proteins. Nature 636, 182–189 (2024). DOI: https://doi.org/10.1038/s41586-024-08220-3
V.J. Hoerschinger, F. Waibl, N.D. Pomarici et al. PEP-Patch: Electrostatics in Protein–Protein Recognition, Specificity, and Antibody Developability. J. Chem. Inf. Model, 63, 22, 6964–6971 (2023). DOI: https://doi.org/10.1021/acs.jcim.3c01490
M.L. Fernández-Quintero, J. Kokot, F. Waibl et al. (2023) Challenges in antibody structure prediction. mAbs, 15(1). DOI: https://doi.org/10.1080/19420862.2023.2175319