April 30, 2020 - by Santina Russo
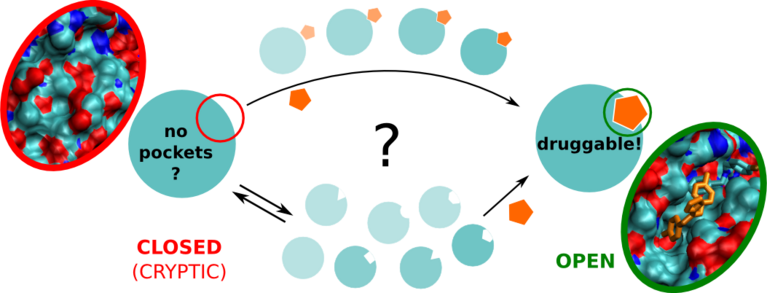
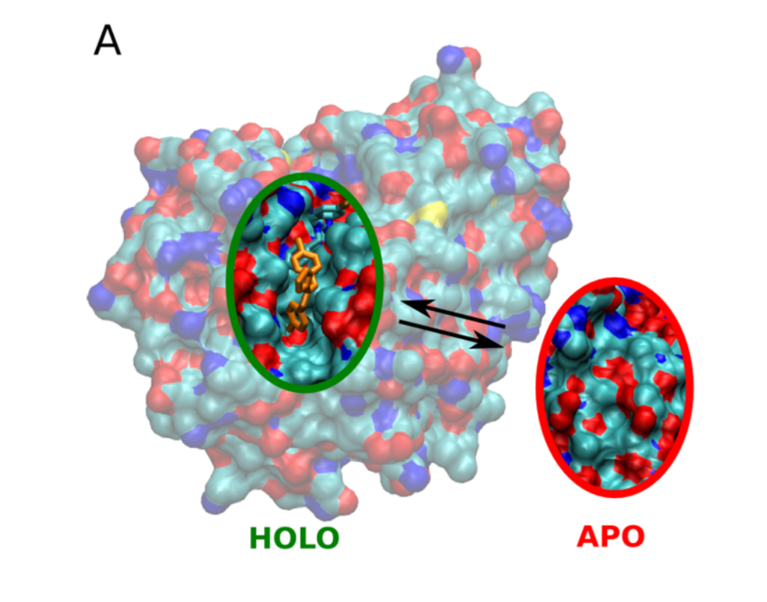
For a long time, some proteins and protein families have been presumed to be undruggable. Even though there was structural information available and a strong medical interest in finding potential drug molecules targeting the proteins, it seemed none could be discovered. “We know now that many of these proteins feature cryptic binding pockets”, says Francesco Gervasio, professor in chemistry and structural biology at the University College London and at the University of Geneva. Cryptic pockets are hidden and appear only when a ligand molecule is actually bound. That’s why, until now, they have only been found by chance and cannot be targeted with conventional drug development methods. However, new findings prove they can be discovered by smart molecular dynamics.
Trick the pockets into opening
Using the “Piz Daint” supercomputer at CSCS, Gervasio and his co-workers developed an enhanced sampling simulation method to discover hidden cryptic binding pockets. Specifically, they used a so-called Hamiltonian replica exchange-based approach for sampling the possible conformational changes of the target proteins. In the process, the hydrophobicity of water is increased, thus increasing the interaction between protein and water. Along with the insertion of various small organic molecules into the system, the combined approach ultimately helps to trick the hidden pockets into opening.
One example where this approach, named SWISH (Sampling Water Interfaces through Scaled Hamiltonians), was successful was the discovery a new binding pocket in the protein K-RAS, an important cancer drug target. K-RAS has a natural ligand that binds so strongly, no drug candidate tested so far could compete with it. But previous experimental results, as well as Gervasio’s method, have found a second cryptic binding pocket — and with this, a second chance to target the protein. Similarly, the SWISH method was able to detect a cryptic pocket in the fibroblast growth factor receptor, another major cancer drug target, and in another protein called TEM1. A so-called beta lactamase, TEM1 causes resistance against antibiotics like ampicillin in bacteria.
Help against the coronavirus
So far, the sampling method has been shown to detect cryptic pockets in proteins reliably, as long as the energy barrier to open the cavities is not too high. In order to study more complex cases, Gervasio is currently improving his algorithm by introducing metadynamics. His next research targets are tumour necrosis factors (TNF), a protein family for which conventional drug design has failed so far.
At the moment though, Gervasio is most occupied with the new SARS-CoV-2 coronavirus, which is currently causing an alarming pandemic around the world. He and his team are using their sampling methods to target Nsp1, a viral protein that has been identified as a potential virulence factor and a target for vaccine development. As previous results on other coronaviruses had shown, Nsp1 plays a crucial role in the downregulation of the host’s immune responses to the virus infection. Interestingly, Nsp1 features no apparent cavities where a ligand could bind, so the search for a cryptic binding pocket could prove decisive in the search for effective treatment.
Understudied protein targets
In addition to these specific examples, the SWISH method allows researchers to further investigate the underlying molecular mechanism of the formation of such cryptic pockets, which is still not quite clear. This new method could even help with investigating whole protein families, like phosphatases. “Phosphatases are important in the formation of cancer, in fact as important as the widely studied kinases”, explains Gervasio. Members of both protein families are involved in the signal pathways that regulate cell proliferation, meaning cell division and thereby generation of new body cells, both for the substitution of damaged or old cells or simply for growth. If elements of these crucial regulatory pathways are malfunctioning, this results in unregulated cell division and, consequently, tumour growth. In fact, similar to kinases, several phosphatases have been shown to promote tumour development and progression.
Unfortunately, phosphatases had been deemed completely undruggable because of previous failures to find ligand molecules that bind to them in a selective way. However, some phosphatases were recently found to have cryptic binding pockets. “Again, their investigation could provide a promising alternative to classical binding sites for drug discovery”, says Gervasio. He is in fact already collaborating with several pharma companies that are interested in using his approach.
This text by Santina Russo first appeared on the PRACE website.
(Image on top: Felipe Esquivel Reed / CC BY-SA)
References:
- Kuzmanic, A. et al.: Investigating Cryptic Binding Sites by Molecular Dynamics Simulations. Acc. Chem. Res. (2020)
- Comitani, F. and Gervasio, F.L.: Exploring Cryptic Pockets Formation in Targets of Pharmaceutical Interest with SWISH. J. Chem. Theory Comput. (2018)
- Oleinikovas, V. et al.: Understanding Cryptic Pocket Formation in Protein Targets by Enhanced Sampling Simulations. J. Am. Chem. Soc. (2016)